If you are interested in the detailed methodology, please refer to our manuscript, BioQueue: a novel pipeline framework to accelerate bioinformatics analysis, for more information.
BioQueue is a researcher-facing platform preferentially to improve the efficiency and robustness of analysis in bioinformatics research. In this post, details about how BioQueue tries to achieve these two goals are explained.
Speeding up analysis
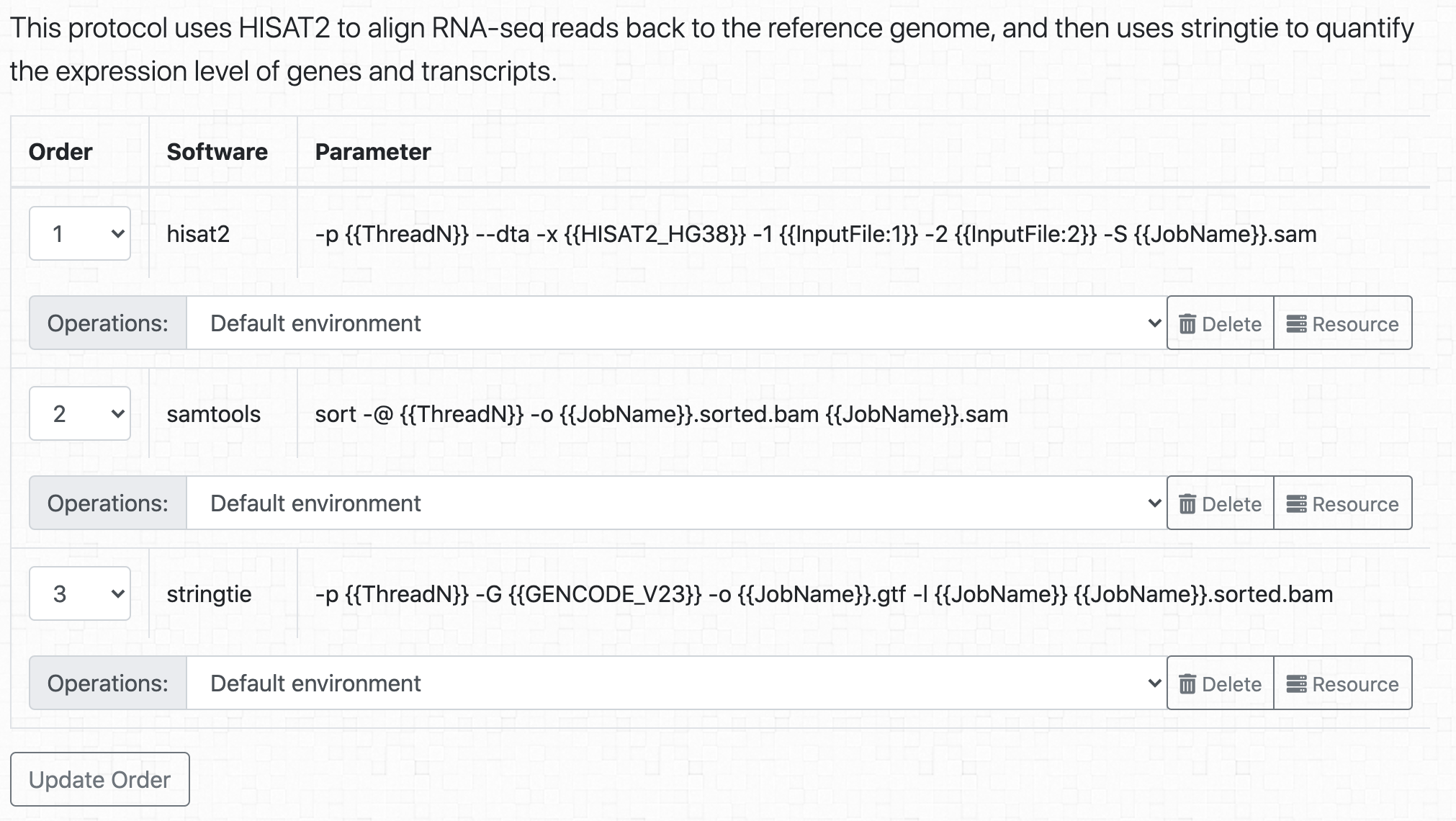
BioQueue organizes analysis workflows as protocols, which are composed of continuous steps. For example, to quantify the expression of genes from RNA-seq samples, we can use the workflow from (Pertea et al., 2016); in BioQueue, it can be expressed as follows:
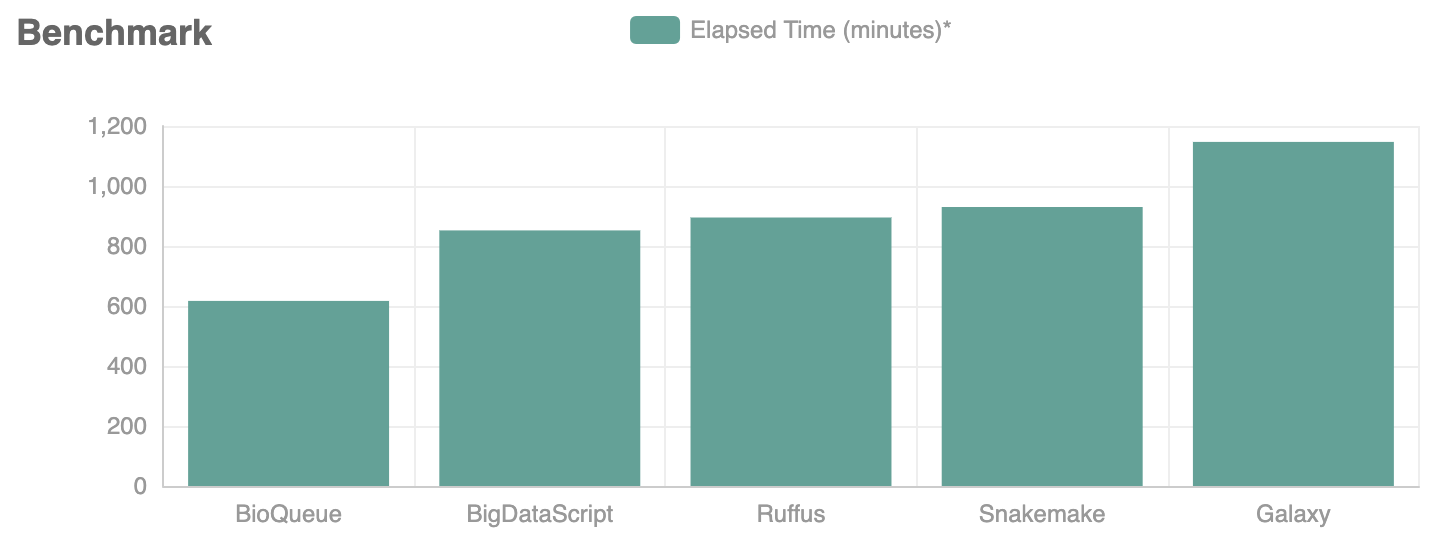
Limited by the design of software or the system resources, many currently available tools cannot fully use the system resources allotted to them or may not achieve excellent efficiency and can even generate errors (such as memory overflow) when running multiple jobs simultaneously. When running each step, BioQueue monitors the actual resources (CPU, peak memory usage, and disk usage) that a step occupies and predicts future resources usage for this step from collected data. It then uses a greedy-algorithm-based dispatcher to arrange the execution order of multiple jobs to ensure the maximum usage of resources and thus speed up the overall efficiency of analysis. For analyzing the same set of data with the identical pipeline, BioQueue can save up to 46% of the required time (boosting rates vary depending on protocols).
Protecting the integrity of results
To protect the integrity of analysis results from human errors (like overwriting a result file by accident), BioQueue actively scans the changes on the inputs and outputs that jobs depend and produce. If files are changed after the job is finished, a color indication (red) will be added to the corresponding job cards.
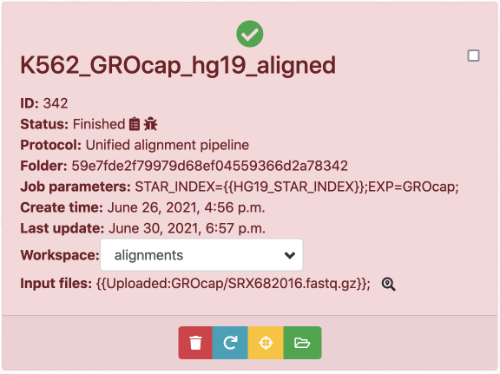
Another common situation is that after a specific period, you may need to rerun the same analysis for different reasons (like you want to test the reproducibility of the results). BioQueue keeps track of the versions of protocols that jobs used, and if a job is not generated with the latest protocol, another color indication (yellow) will be added to the job cards warning that rerun the job may yield inconsistent results.
BioQueue also supports achieving job files and backup them to a different destination to prevent the effect of disk failure.
Other handy functions
- Steps can be executed in different conda/venv environments so that you can handle dependencies that different tools/software requires with ease;
- BioQueue provides a registry of biosamples, and automatically associates jobs with registered samples;
- BioQueue has two types of account groups: worker (with read and write permission) and viewer (read only). You can create viewer accounts and share the account info with your partners, they will be able to download files, and you don’t need to worry about jobs’ safety.
Useful links: